Dr Christos Pliotas
- Position
- Lecturer in Integrative Membrane Biology
- Areas of expertise
- Mechanosensitive channels; EPR spectroscopy; PELDOR/DEER; Membrane proteins; CryoEM;
- Phone
- +44(0)113 343 1229
- Location
- Astbury 6.108a
- Faculty
- Biological Sciences
- School
- BiomedicaL Sciences
Introduction
Membrane pores sense and respond to membrane forces, in order to regulate the flux of ions, which is essential for cell signalling and physiology. Ligands that would enable capturing the distinct states these systems undergo during opening and closure (a process also known as gating) are not known, thus their investigation by any method is challenging.
We study mechanosensitive ion channels by PELDOR/DEER spectroscopy and CryoEM, in order to understand the fundamental biophysical property of mechanosensation.
Mechanosensitive channels are ubiquitous across life kingdoms and in bacteria they protect the cell against severe osmotic shock, by acting as pressure valves to relieve cytoplasmic pressure. In humans, mechanosensitive channels are involved in hearing, touch and cardiovascular architecture. Key to their regulation is the lipid, the building block of membranes. It is the lipid’s dynamic nature which allows the transmission of force and curvature deformation in the membrane to directly activate mechanosensitive ion channels. The aim of our group is to dissect the molecular basis of mechanosensation and elucidate the role of lipids in the regulation of ion channels and other force-sensitive membrane proteins.
Current major projects
- Allosteric activation of mechanosensitive ion channels
- PELDOR/DEER spectroscopy and spin labelling of membrane proteins
- CryoEM of ion channels
- Functional multimodality
Detailed research programme
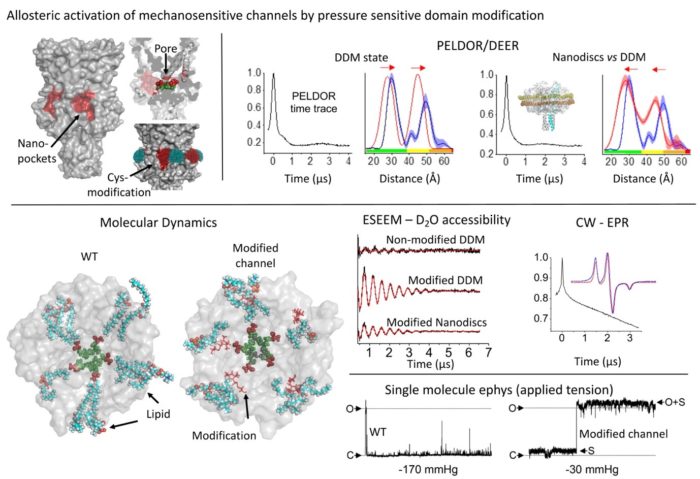
Allosteric activation of mechanosensitive ion channels
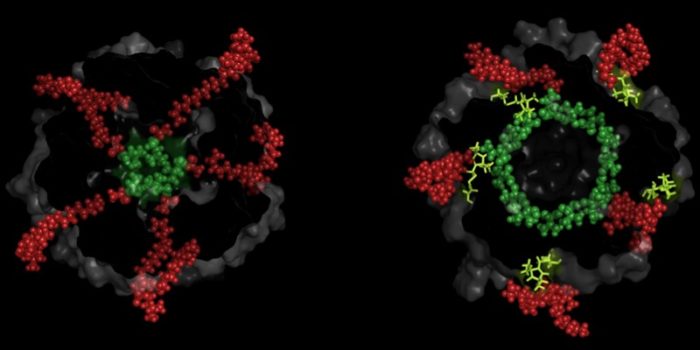
Lipid availability within hydrophobic nano-pockets has been linked with mechanosensation in ion channels, across life. However, the effect of hindering lipid access into these nano-pockets on channel structure is not known. We identified such nano-pockets on the MscL mechanosensitive channel, the highest pressure activation threshold channel in nature. We restricted lipid-chain access by cysteine modification, and interrogated MscL’s conformation by PELDOR/DEER spectroscopy. For a site located at the nano-pocket entrance and distal to the channel pore we generated an allosteric response in the absence of applied membrane force. This response was fully reversed following incubation with a reducing agent. The modification associated with the conformational change, interrupted the contact between the membrane and the channel which mediated mechanosensitivity.
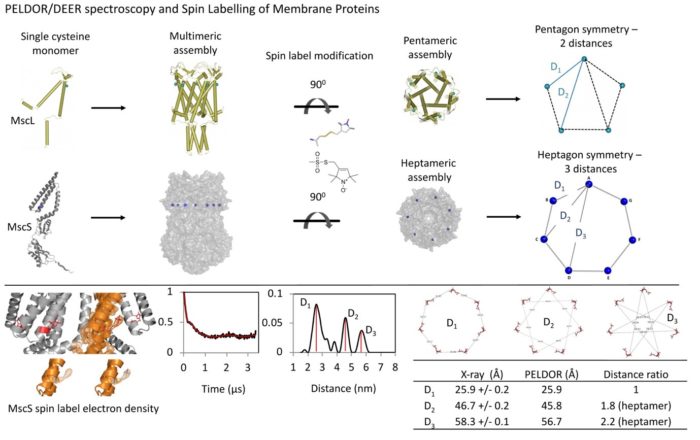
EPR (PELDOR/DEER) spectroscopy for the investigation of membrane protein folding, structure and dynamics
Multimeric (or monomeric with two or more cysteines per subunit) proteins carry more than one spin label and pulse electron paramagnetic resonance (EPR), PELDOR/DEER spectroscopy is facilitated to obtain spin-to-spin distances within a protein complex. These distances, could be used as restraints to resolve the protein’s dynamic and oligomeric state(s), under physiological conditions and within native/lipid environment. Alongside PELDOR, Electron Spin Echo Envelope Modulation (ESEEM) spectroscopy could be used to measure deuterium (solvent) accessibility of key residues, important in protein’s function.
PELDOR and ESEEM could also constitute main pipeline methods along with Cryo-Electron Microscopy (CryoEM) and Hydrogen Deuterium Exchange Mass Spectrometry (HDX-MS) to facilitate 3D structure determination of membrane proteins.