Professor David Brockwell
- Position
- Professor
- Areas of expertise
- Proteins; Force; Flow; Aggregation; Biopharmaceuticals
- Phone
- +44(0) 113 343 7821
- Location
- Astbury 10.116
- Faculty
- Biological Sciences
- School
- Molecular and Cellular Biology
Introduction
Proteins carry out a bewildering array of diverse functions that are essential for life and which are increasingly being exploited by industry. The efficiency and specificity at the heart of protein function is often due to their inherent dynamism and relative instability. Consequently, small changes in the environment or the presence of perturbants such as temperature or mechanical force can lead to their unfolding and aggregation. The Brockwell group uses a range of biophysical and protein engineering methods to understand the relationships between sequence, structure, stability, function and aggregation for disease-relevant and biopharmaceutical proteins.
Current major projects
- Identification and amelioration of aggregation-prone biopharmaceuticals
- Understanding effects of mechanical force and hydrodynamic flow on proteins
- Outer membrane protein folding in vitro and in vivo
- Design, construction and characterisation of protein hydrogels
Detailed research programme
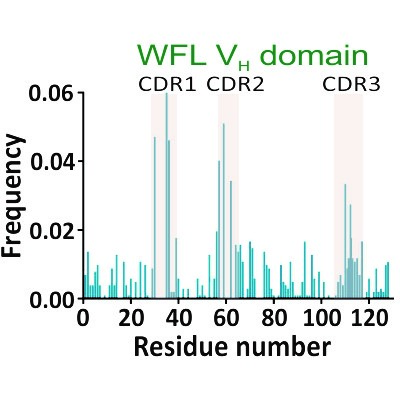
Biopharmaceutical manufacture
The aggregation of metastable proteins is highly problematic in the biopharmaceutical industry where hydrodynamic forces encountered during manufacture can trigger aggregation, causing a loss of efficacy and adverse immune responses upon administration.
Identifying which of the many candidate sequences are inherently robust to manufacturing is difficult. In a collaboration with Professors Nik Kapur (School of Mechanical Engineering) and Sheena Radford, we are investigating the mechanism of flow-induced aggregation (Dobson et al. 2017) Proc. Natl. Acad. Sci. USA), the utility of bench-top hydrodynamic flow devices to assess manufacturability (Willis et al. Eng Rep (2020)) and the use of an in vivo screen to assess and modulate (using a directed evolution approach) the ‘manufacturability’ of biopharmaceuticals (Ebo et al. (2020) Nature Commun).
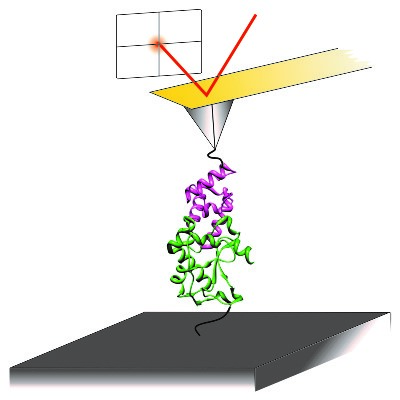
The effects of force on proteins and their complexes
Many proteins resist or respond to mechanical stimuli. We use atomic force microscope (AFM) instruments to investigate the effects of force on single proteins and their complexes. This work has revealed that proteins can behave very differently under force compared to more traditional denaturants (Sadler et al. (2009) J Mol Biol). For example, some extremely thermodynamically stable complexes are exquisitely sensitive to force (Farrance et al. (2013) PLoS Biol) yet other moderately avid complexes are strong enough to mechanically gate outer membrane transporters (Hickman et al. (2017) Nature Commun). We are currently investigating the origins of the extremely high mechanical strength of SasG – an extracellular S. aureus protein that facilitates colonisation and biofilm formation (Gruszka et al. (2015) Nature Commun).
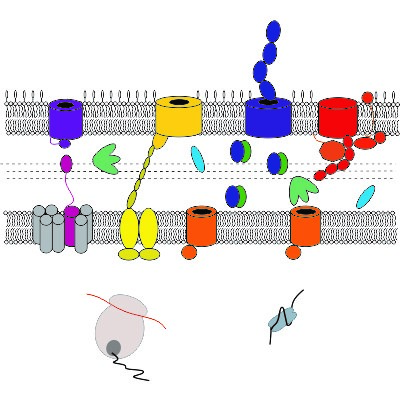
Membrane protein folding and folding factors
The cell envelope is essential for Gram-negative bacteria and is packed with outer membrane proteins (OMPs). Despite their importance as virulence factors, understanding how OMPs fold to their functional form is poor. In collaboration with Professor Sheena Radford we have previously delineated the folding mechanism of a bacterial OMP in vitro (Huysmans et al. (2010) Proc Natl Acad Sci USA and Huysmans et al. (2012) J Mol Biol). Current research is centred on the role of periplasmic chaperones (Schiffrin et al. (2016) Nature Struct. Mol. Biol. and Calabrese et al. (2020) Nature Commun) and (with Professors Neil Ranson and Sheena Radford) how the essential BAM complex facilitates this process (Schiffrin et al. (2017) J Mol Biol and Iadanza et al. Nature Commun (2016)).
Protein hydrogels
Protein hydrogels are macroscopic three dimensional, hydrated, highly porous, percolating networks that have found applications in tissue engineering and are being developed as ligand-triggered actuators for biosensors and for controlled release for drug delivery.
Currently as most protein-based hydrogels are obtained from unstructured peptides or through aggregation of unfolded globular proteins, the full spectrum of protein function (e.g., catalysis, signalling, and ligand binding) has not yet been exploited. In a collaboration with Professor Lorna Dougan (School of Physics and Astronomy) we are building hydrogels from tandem arrayed, folded globular proteins with known mechanical properties assessed using AFM and investigating the emergent macroscopic rheological properties (Hughes et al. (2020) Soft Matter).