Dr Juan Fontana
- Position
- Associate Professor
- Areas of expertise
- virus structure; electron microscopy; cryo-EM; cryo-ET; cellular EM
Introduction
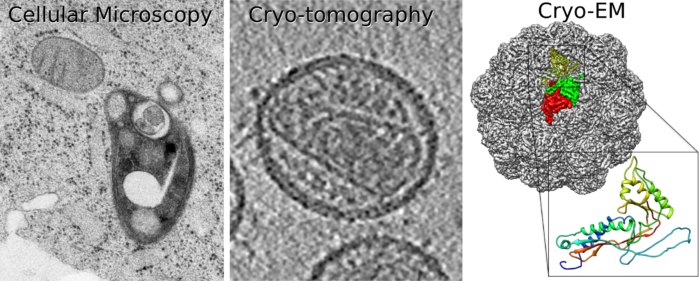
Research in my lab is focused on the structural characterisation of macromolecular assemblies, and viruses that are a current public health risk (like HIV, Influenza, herpesviruses and bunyaviruses), using cryo-EM, combined with other structural approaches.
Current major projects
- Structural biology of enveloped viruses
- Influenza virus entry and assembly
- Ribosomal translational regulation
- Electron microscopy of macromolecular assemblies
Detailed research programme
Structural biology of enveloped viruses
One of the main areas of research of the lab is the structure of enveloped viruses. For example, we have studied the requirement of endosomal ions for enveloped virus fusion and found that K+ ions induce conformational changes in the viral fusion proteins of bunyaviruses, potentially enhancing interactions with the target membrane. We have also characterised the pre-fusion conformation of Herpes Simplex Virus fusion protein, and shown that its fusion loops point towards the viral membrane.
Influenza virus entry and assembly
Influenza virus is a current public health risk. 5-10% of adults are yearly affected by seasonal Influenza outbreaks, leading to up to 5 million cases of severe illness and 500,000 deaths worldwide. Additionally, Influenza can develop resistance to current antivirals and new strains can emerge and result in pandemics. Our lab is interested in two aspects of influenza virus: 1) how influenza starts an infection within a cell, a process that requires the merging (fusion) of the viral and cellular membranes mediated by the viral hemagglutinin protein; and 2) how influenza’s 8 RNA segments bundle together to form an infectious particle, potentially resulting in novel pandemic influenza viruses. We are also looking into developing strategies to detect and block influenza virus infections.
Ribosomal translational regulation
We are interested in understanding the translational control exerted by ribosomes. For example, we are currently studying how paralog switching in Drosophila melanogaster ribosomes leads to translation regulation. By purifying ribosomes from different Drosophila tissues, we have found that Drosophila’s ribosomes are heterogeneous, mainly via paralog switching, and we have found structural differences between ribosomes from testes and ovaries. These differences are compatible with translation regulation by specialised ribosomes.
Electron microscopy of macromolecular assemblies
Electron microscopy (EM) allows the study of macromolecules at atomic to molecular resolution. We take advantage of our expertise in cryo-EM and cryo-electron tomography to study a number of macromolecule assemblies. For example, we are interested in the ultrastructure of the extracellular matrix, which is involved in a variety of processes, from fertility to inflammation.
