Dr Neil Thomson
- Position
- Reader in Biological Physics and Bionanotechnology
- Areas of expertise
- Bionanotechnology; Biological Physics; synthetic biology; atomic force microscopy; DNA
- Phone
- +44(0)113 343 7289
- Location
- GR24C Sir William Henry Bragg Building
- Faculty
- Engineering and Physical Sciences
- School
- Physics and Astronomy
Introduction
My group develops and applies atomic force microscopy (AFM) techniques to study the structure, dynamics and interactions of nanoscale systems, down to the molecular level. These systems are typically self-assembled and can include inorganic, biomolecular and/or hybrid materials. AFM is a versatile technique that we use to image, measure forces or manipulate material. We carry out high resolution imaging either in situ in aqueous liquid environments or ex situ in air, depending on the application. We also make force measurements to determine the mechanics of materials and molecules at the nanoscale. All work is collaborative with biological scientists and other physical scientists creating a multi-disciplinary team and working environment. Where appropriate, complementary biochemical and biophysical techniques are used to enhance understanding gained from the AFM data.
A key goal is to gain understanding of complex systems down to the molecular scale, aiming to develop new materials and applications with focus primarily on healthcare. We are particularly interested in DNA systems, both as a synthetic nanomaterial and as the genetic component of life. Collaborations at Leeds include hydrogels for tissue engineering with colleagues in the Dental School and therapeutic microbubbles for targeted cancer treatment as part of the Leeds Microbubble Consortium.
Current major projects
- Concurrent DNA transcription in simple gene models
- DNA origami: structure and self-assembly
- Self-assembly of nanoscale systems at liquid-liquid interfaces
Detailed research programme
Concurrent Transcription
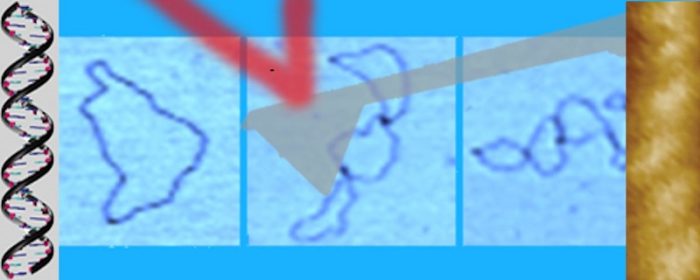
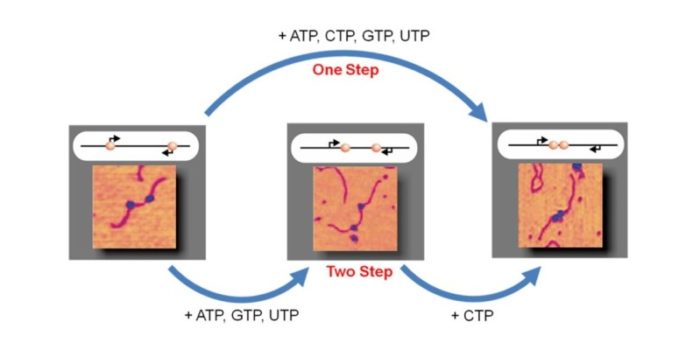
The organisation of genes within a genome is complex and may have regulatory functions arising from evolutionary pressure over eons of time. Genetic structures are increasingly being discovered in compressed structures, such as nested within one another or with closely spaced promoters but divergently aligned. We use single molecule AFM imaging to visualise outcomes of RNA polymerase traffic on DNA templates with two (or more) genes, either in convergent or tandem configurations. The efficiency of concurrent transcription is influenced by the asynchronous behaviour of multiple polymerases acting on the DNA. This brings new understanding to the fundamental understanding of DNA transcription and has implications for gene expression, developmental biology and genetic conditions underlying disease states.
DNA origami
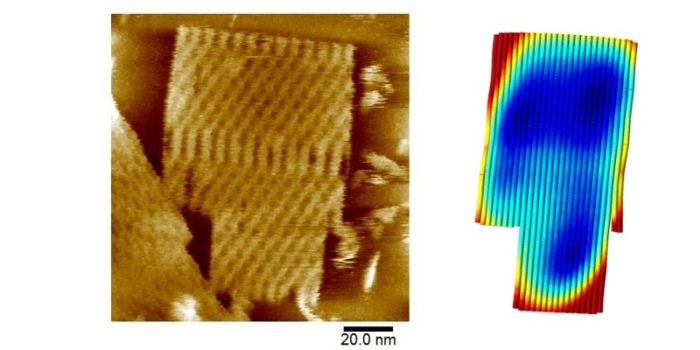
DNA origami is a developing and potentially underpinning biomolecular nanotechnology that uses rational design of short DNA oligomers to self-assemble or fold a longer single-stranded DNA template into complex 2D and 3D patterns and objects on the order of 100nm in size. A long-term goal of this field is to develop nanoscale molecular devices with high efficiency to carry out tasks such as energy harvesting and transduction, computation and therapeutic delivery for healthcare applications. Our group is studying the structure and dynamics of single sheet DNA origami nanotiles using high resolution atomic force microscopy. Understanding in greater detail how to control origami assembly and structure is key to developing new materials applications based on DNA nanotechnology.
Liquid-Liquid Interfaces
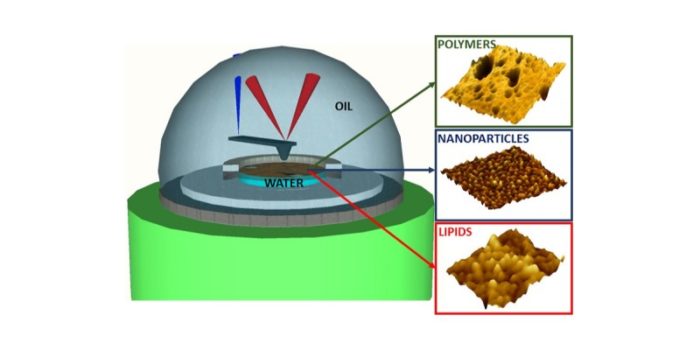
Atomic force microscopy is typically performed at the solid-liquid or solid-air interface both to gain high resolution surface profiles of samples or determine the nanomechanics of surfaces and/or molecules. Many nano systems, including inorganic and biomolecular, self-assemble at liquid-liquid interfaces. Our recent collaboration with the European Synchrotron Radiation Facility (ESRF) in Grenoble, France, achieved the first nanoscale images of nanoparticles, lipids and polymers assembled at liquid-liquid interfaces using AFM. Grazing incidence small angle X-ray scattering data from ESRF correlated perfectly with the real-space AFM data for the self-assembled structure of silicon nanoparticles at oil/water interfaces. This work is being expanded to investigate more challenging biomolecular systems at the liquid-liquid interface.