Dr Robin Bon
- Position
- Associate Professor of Chemical Biology
- Areas of expertise
- TRPC cation channels; chemical protein labelling; photoaffinity labelling; mass spectrometry; cryo-EM
- r.bon@leeds.ac.uk
- Phone
- +44(0) 113 343 0785
- Location
- Level 7 LIGHT Laboratories
- Faculty
- Medicine and Health
- School
- Leeds Institute of Cardiovascular and Metabolic Medicine, School of Medicine
Introduction
We use chemical, biochemical and biophysical approaches to understand the molecular mechanisms underlying health/disease and the effects of bioactive small molecules. For example, we use site-directed mutagenesis, cellular assays, new photoaffinity probes, mass spectrometry and cryo-electron microscopy to develop a detailed understanding of interactions between bioactive small molecules and proteins (e.g., TRPC1/4/5 cation channels and the mitochondrial chaperonin HSPD1). In addition, we develop novel chemical tools for biomolecule labelling and mass spectrometry to analyse biological samples. The group has lab space in the LIGHT laboratories (mammalian cell culture, biochemistry, pharmacology) and in the School of Chemistry (organic and analytical chemistry) and collaborates with several Astbury colleagues (e.g., biological mass spectrometry, cryo-EM).
Current major projects include
- Chemical and structural biology of TRP cation channels
- The mitochondrial chaperonin HSPD1 as a potential therapeutic target in glioblastoma
- LALDI mass spectrometry for analysis of biological samples
Detailed research programme
Chemical and structural biology of TRP cation channels
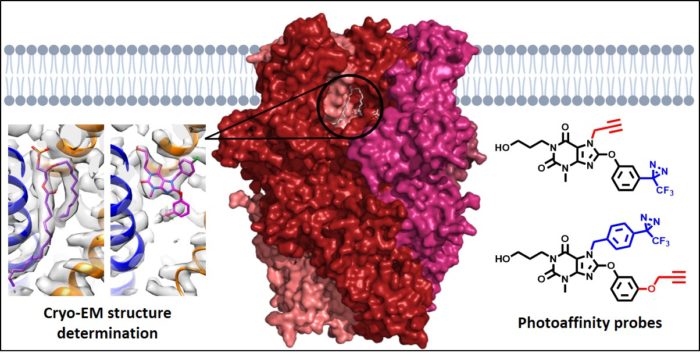
Transient Receptor Potential (TRP) proteins, which include the TRPM, TRPV, TRPA and TRPC subtypes, form tetrameric, non-selective cation channels permeable by Na+ and Ca2+. For all 28 mammalian TRP proteins, four monomers are needed to form a functional ion channel, and channels may consist of homomers or heteromers of subunits, each with their own characteristics and functions. This project focuses on homomeric and heteromeric channels formed by TRPC1, TRPC4 and TRPC5. Their implication in various human disease states has led these channels to emerge as potential therapeutic targets. However, little is known about the exact composition of TRPC1/4/5 tetramers in different tissues, and the lack of information about binding modes of channel modulators prevents the design of tetramer-specific activators and inhibitors. In this project, we use molecular biology, photoaffinity probes, mass spectrometry and cryo-electron microscopy to unravel the interactions between specific TRPC1/4/5 subunits, and the interactions between TRPC1/4/5 channels and small-molecule modulators, lipids and metal ions.
The mitochondrial chaperonin HSPD1 as a potential therapeutic target in glioblastoma
Gliomas are cancerous brain tumours that aggressively invade and grow within healthy brain. Across the world, 250,000 people are diagnosed with brain cancer every year, and this devastating disease accounts for 189,000 deaths per year (UK: 3,500 deaths p.a.). Glioblastoma multiforme (GBM) is the most common and aggressive form of brain cancer. Even with the best treatment, GBM patients only live for 12-15 months on average, and fewer than 1 in 20 patients live more than 5 years. This is mainly because some GBM cells that cannot be safely removed by surgery (without affecting normal brain function) are resistant to current chemo- and radiotherapy, leading to regrowth of brain tumours. The small molecule KHS101 selectively kills a wide range of patient-derived GBM cells, but not healthy brain cells, in vitro and in mice, by inducing a catastrophic metabolic phenotype. Through a major interdisciplinary collaboration that involved chemical proteomics, global proteomics, transcriptomics, metabolomics, and assays in vitro, in cells and in animal tissues, the mitochondrial chaperonin HSPD1 was identified and validated as the relevant target of KHS101 in GBM cells. We are currently using a combination of photoaffinity labelling, mass spectrometry and cryo-electron microscopy to develop a deeper understanding of the molecular interactions between HSPD1 and KHS101, in order to enable structure-based design of new HSPD1 inhibitors for treatment of GBM.
LALDI mass spectrometry for analysis of biological samples
Label-assisted laser desorption/ionisation mass spectrometry (LALDI-MS) is a novel and powerful tool for the selective analysis of labelled species by mass spectrometry. LALDI-MS is carried out in a similar manner to matrix-assisted laser desorption/ionisation mass spectroscopy (MALDI-MS), however the external matrix component of MALDI has been replaced with a laser desorption/ionisation-enhancing label. Through this development, LALDI-MS has gained several benefits over MALDI-MS, eradicating concerns over optimum analyte–matrix pairings and allowing analysis to be performed directly against a complex background without the need for purification. We develop novel, water-soluble, LALDI tags for in situ labelling of analytes in complex samples. The technology allows us to directly analyse biological samples without the need for purification.