Professor Paul Taylor
- Position
- Professor of Chemical Education
- Areas of expertise
- Molecular Evolution; Chemical Genomics; Cancer
- Phone
- +44(0)113 343 6529
- Location
- G.52c Chemistry
- Faculty
- Engineering and Physical Sciences
- School
- Chemistry
- Website
- ORCID
Introduction
It is widely accepted that eukaryogenesis involved an endosymbiosis between at least one archaeon, probably from the Asgard lineage, and at least one bacterium, most likely an ancestral -proteobacterium. More controversial and less well understood is the possibility of further symbioses or other significant gene flows from bacteria to ancestral eukaryotes. Both the order in which these key evolutionary events occurred and their timeline are unclear. Characterising the origins of animals from this ancient mix is of particular interest.
We use phylogenetic analyses to explore the evolution of chemical signalling processes and how the characteristic signalling pathways in animals emerged during eukaryogenesis. Current areas of interest are retinoid signalling, calcium signalling and peroxynitrite signalling.
Our more applied research uses chemical genomics approaches to find new cancer relevant targets for existing drugs and explores how dietary nutrients interact with cancer treatments, as exemplified by VAL401, an adenocarcinoma treatment currently in clinical trials for non-small cell lung cancer.
Current major projects
- The bacterial origins of chemical signalling in animals
- Chemical genomics approaches to drug reprofiling for cancer treatment
Detailed research programme
The Bacterial Origins of Chemical Signalling in Animals
Major advances in understanding of human biology have been made by studying so-called "model organisms", such as mice, fruit-flies and yeast that are readily handled under laboratory conditions. Nevertheless, there are still large gaps in our knowledge. The genomics revolution has made available the sequenced genomes of thousands of diverse organisms meaning scientists can now potentially study healthy systems or disease states of interest in entirely different species.
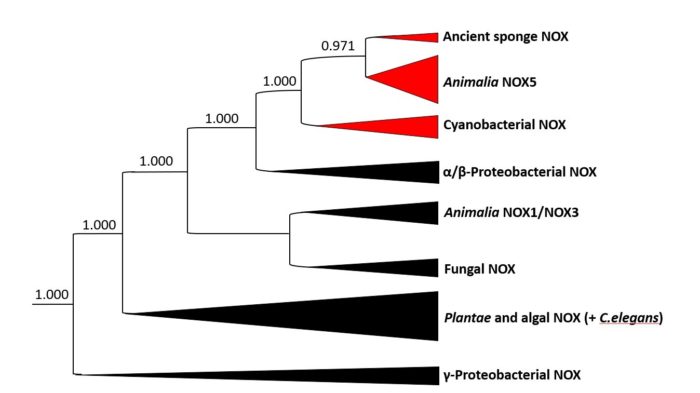
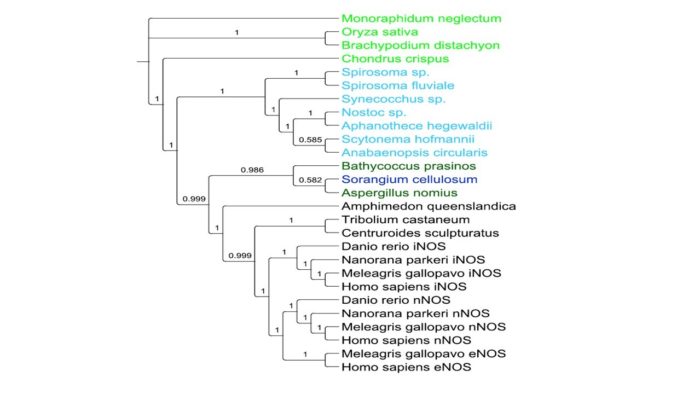
Phylogenetic analysis and experimental results from labs around the world, including ours, indicate the presence of animal-like chemical signalling systems in species of cyanobacteria. Through this programme, we will consolidate these phylogenetic, chemical, biochemical and microbiological observations to offer new perspectives on evolution and opportunities for innovative research in the biomedical sciences.
Chemical Genomics Approaches to Drug Reprofiling for Cancer Treatment
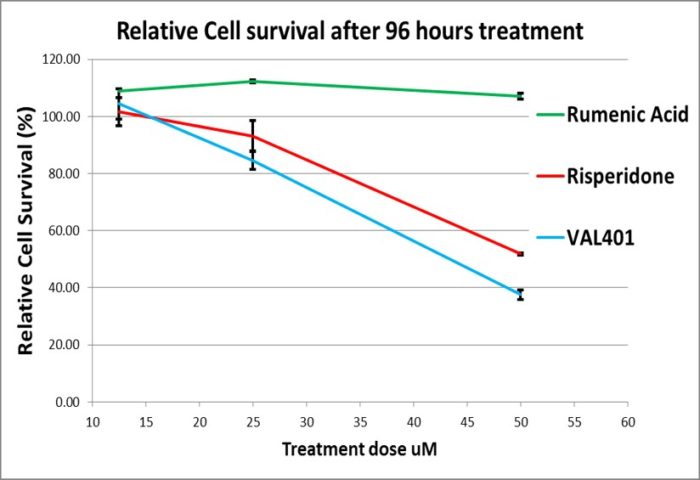
Discovered using the Magic-Tag chemical genomics platform, VAL401 is a reformulation of risperidone, an antipsychotic used since 1993 as a first-line treatment for schizophrenia and bipolar disorder. We found that risperidone inhibits HSD10 (hydroxysteroid dehydrogenase type 10), an enzyme involved in the processing of steroids and fats. Normally located in mitochondria, HSD10 is overexpressed in cancer cells, where it is also accessible from the cytoplasm. Inhibition of HSD10 disrupts cancer energy metabolism, breaking the cancer cell cycle.
Although risperidone has no anticancer effects when administered alone, its anticancer activity is switched on when it is combined with rumenic acid, a nutritional supplement derived from safflower oil.
We continue to study the science underlying the action of VAL401.