Professor Peter Henderson
- Position
- Emeritus Professor
- Areas of expertise
- Structures and mechanisms of membrane transport proteins; resistance of bacteria to antimicrobials; applications of microbial efflux transport in Biotechnology
- Website
- ORCID
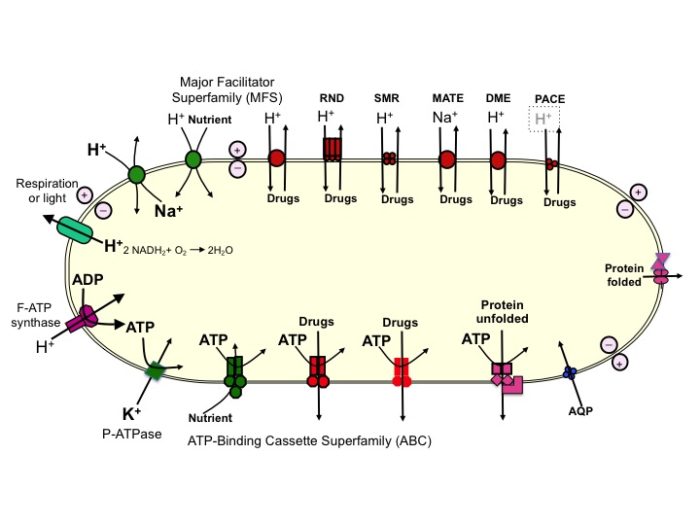 We aim to elucidate the structure-activity relationships of a wide range of bacterial membrane transport proteins. The focus is on efflux proteins that confer resistance to antimicrobials in a number of important pathogens such as Acinetobacter baumannii, Pseudomonas aeruginosa, Vibrio haemolyticus, Bordetella pertussis, Helicobacter pylori, Brucella melitensis and others. Our strategy is to transfer genes encoding transport proteins to suitable vectors for amplification of expression in Escherichia coli and subsequent purification for structural and mechanistic studies. A wide range of techniques are exploited to characterise the proteins – X-ray crystallography, MS, HDX, NMR, fluorimetry, CD and FTIR, EPR spectroscopy and Molecular Dynamics Simulations. Activities are assayed by measuring transport of radioisotope-labelled substrates into cells, or into proteoliposomes containing purified proteins, and/or by measurements of unlabelled ligand binding to the proteins. Fluorescent indicators are particularly useful. Mutants and/or chimaeric proteins are also examined to locate substrate/inhibitor binding sites, determine recognition of cations, define steps in the translocation process, facilitate structural studies, understand conformational triggers, etc. Inhibitors of efflux are being developed to help overcome Antimicrobial Resistance (AMR). Organisms specially engineered to synthesise feedstocks for commercial exploitation are genetically modified to enhance efflux of desired, and possibly toxic, products.
We aim to elucidate the structure-activity relationships of a wide range of bacterial membrane transport proteins. The focus is on efflux proteins that confer resistance to antimicrobials in a number of important pathogens such as Acinetobacter baumannii, Pseudomonas aeruginosa, Vibrio haemolyticus, Bordetella pertussis, Helicobacter pylori, Brucella melitensis and others. Our strategy is to transfer genes encoding transport proteins to suitable vectors for amplification of expression in Escherichia coli and subsequent purification for structural and mechanistic studies. A wide range of techniques are exploited to characterise the proteins – X-ray crystallography, MS, HDX, NMR, fluorimetry, CD and FTIR, EPR spectroscopy and Molecular Dynamics Simulations. Activities are assayed by measuring transport of radioisotope-labelled substrates into cells, or into proteoliposomes containing purified proteins, and/or by measurements of unlabelled ligand binding to the proteins. Fluorescent indicators are particularly useful. Mutants and/or chimaeric proteins are also examined to locate substrate/inhibitor binding sites, determine recognition of cations, define steps in the translocation process, facilitate structural studies, understand conformational triggers, etc. Inhibitors of efflux are being developed to help overcome Antimicrobial Resistance (AMR). Organisms specially engineered to synthesise feedstocks for commercial exploitation are genetically modified to enhance efflux of desired, and possibly toxic, products.
